The 2019 Nobel Prize in physiology or medicine has been awarded to three scientists, William G. Kaelin Jr. from the Dana-Farber Cancer Institute in Boston, Sir Peter J. Ratcliffe from the Francis Crick Institute in London and Gregg L. Semenza from Johns Hopkins University in Baltimore “for their discoveries of how cells sense and adapt to oxygen availability”.
Identifying the molecular machinery that regulates the activity of genes in response to varying levels of oxygen has been key to understanding human diseases such as cancer, anaemia and many others.
Oxygen is paramount to almost all life. It is used by almost all animal cells to convert food into energy. Adaptation to changes in oxygen levels is one of the most important adaptive systems for animal life, according to Randall Johnson, a member of the Nobel Prize Committee.
“Just as a candle needs the right amount of oxygen to burn cleanly, cells need to adjust their metabolic rates based on how much oxygen they have available,” said Johnson in Inside Science. “This allows each cell and indeed our bodies too efficiently and safely burn fuel so as to create heat to do work and build new tissues.”
The fundamental importance of oxygen had been understood for centuries, but how cells adapt to changes in levels of oxygen had long been unknown. A rise in the hormone erythropoietin (EPO) linked to the increased production of red blood cells had been known as a key physiological response to hypoxia since the beginning of the 20th century, but how this process is itself controlled by O2 remained a mystery.
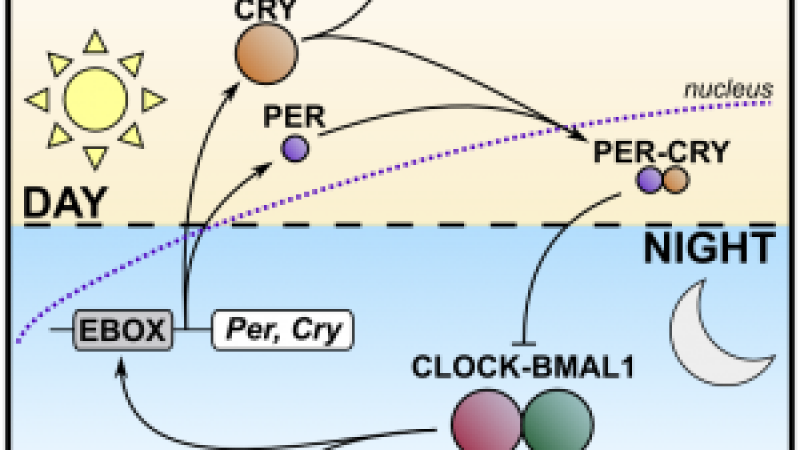
Gregg Semenza studied the EPO gene and how it is regulated by varying oxygen levels using in vivo, in vitro and patient studies. In his efforts, he discovered the hypoxia-inducible factor (HIF) protein complex partly responsible for mediating this response.
“By eliminating the expression of HIF in mice, he was able to uncover the different pathways involved in hypoxia” explains Nathalie Mazure, CNRS researcher at the Université de Nice/Inserm in France. “This couldn’t have been possible in humans and the use of cells wouldn’t have pictured the effect across the whole organism.”
“He was the first to create a knock out mouse model for a gene responding to hypoxia, HIF1,” adds Carole Peyssonauxan Inserm researcher at the Cochin Institute in Paris.
“Semanza’s work in mice was essential, and he couldn’t have completed his work without the use of animals. The mouse models helped show how essential this transcription factor is. It was the only way to find out how the protein works in a tissue specific manner with a whole organism perspective. There is no other way to show the importance of a gene in physiological conditions than to work in vivo.”
Together with Peter Ratcliffe, Semenza found that the oxygen sensing mechanism was present in virtually all tissues, not only in the kidney cells where EPO is normally produced.
Around the same time, cancer researcher William G. Kaelin was looking at an inherited disease called von Hippel-Lindau’s syndrome (VHL disease) which was associated with a higher risk of cancer. He found that the VHL gene - responsible for the disease – could prevent the onset of cancer and was also involved in controlling hypoxia-regulated genes in the cell. Ratcliffe then showed that VHL interacted with HIF-1α and influenced oxygen levels. In 2001, Kaelin and Ratcliffe completed the story by simultaneously showing how that process worked.
Since then, more than 300 genes have been found to be sensitive to oxygen levels involved in a variety of physiological processes and a large number of diseases. The work of all three researchers has had direct therapeutic applications, in particular in cancer research as the generation of a hypoxic environment and activation of its main effector HIF-1, are common features of advanced cancers. « Dissecting the different mechanisms involved in hypoxia, has, every time, given access to new therapeutic targets,” concludes Nathalie Mazure. And there is still much more to uncover.
Key publications
Semenza, G.L, Nejfelt, M.K., Chi, S.M. & Antonarakis, S.E. (1991). Hypoxia-inducible nuclear factors bind to an enhancer element located 3’ to the human erythropoietin gene. Proc Natl Acad Sci USA, 88, 5680-5684
Wang, G.L., Jiang, B.-H., Rue, E.A. & Semenza, G.L. (1995). Hypoxia-inducible factor 1 is a basic-helix-loop-helix-PAS heterodimer regulated by cellular O2 tension. Proc Natl Acad Sci USA, 92, 5510-5514
Maxwell, P.H., Wiesener, M.S., Chang, G.-W., Clifford, S.C., Vaux, E.C., Cockman, M.E., Wykoff, C.C., Pugh, C.W., Maher, E.R. & Ratcliffe, P.J. (1999). The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature, 399, 271-275
Mircea, I., Kondo, K., Yang, H., Kim, W., Valiando, J., Ohh, M., Salic, A., Asara, J.M., Lane, W.S. & Kaelin Jr., W.G. (2001) HIFa targeted for VHL-mediated destruction by proline hydroxylation: Implications for O2 sensing. Science, 292, 464-468
Jakkola, P., Mole, D.R., Tian, Y.-M., Wilson, M.I., Gielbert, J., Gaskell, S.J., von Kriegsheim, A., Heberstreit, H.F., Mukherji, M., Schofield, C.J., Maxwell, P.H., Pugh, C.W. & Ratcliffe, P.J. (2001). Targeting of HIF-α to the von Hippel-Lindau ubiquitylation complex by O2-regulated prolyl hydroxylation. Science, 292, 468-472
https://theoncologist.alphamedpress.org/content/9/suppl_5/10.full
https://www.nobelprize.org/prizes/medicine/2019/press-release/
https://www.insidescience.org/news/2019-nobel-prize-medicine-story
https://www.nature.com/articles/d41586-019-02963-0
Last edited: 4 March 2022 08:57